There are two ways to use RNA-Pareto. The first is an interactive graphical user interface written in Java. The second is a command-line tool, which allows to process the calculated solutions further in own programs.
To start the GUI, click on "RNAPareto.sh" (on Linux), "RNAPareto.exe" (on Windows) or on the RNAPareto app (on Mac), or just type "java -jar RNAPareto.jar" on the command line. An RNA database in FASTA format can then be loaded. After choosing two RNA sequence from this databse, click "Start" to begin the calculation of the Pareto-set of solutions. Using the scatter plot of solutions in the center of the window, choose interesting solutions, which are then displayed.
There is also a standalone command-line tool, which is called "rna-pareto" (on Linux or Mac) or "rna-pareto.exe" (on Windows). This program outputs all Pareto-optimal solutions, each consisting of 5 lines:
The C++ part can be compiled from source. Therefore, the Vienna RNA library has to be present, as well as some Java JDK. There are three Makefiles, because they differ for Linux, Mac and Windows. You may have to change some paths in the makefiles to get all to compile. The Java part was developed using Eclipse IDE. Therefore it should be possible to import the project into eclipse. To get it to work, the appropriate JNI library has to be present in the working directory.
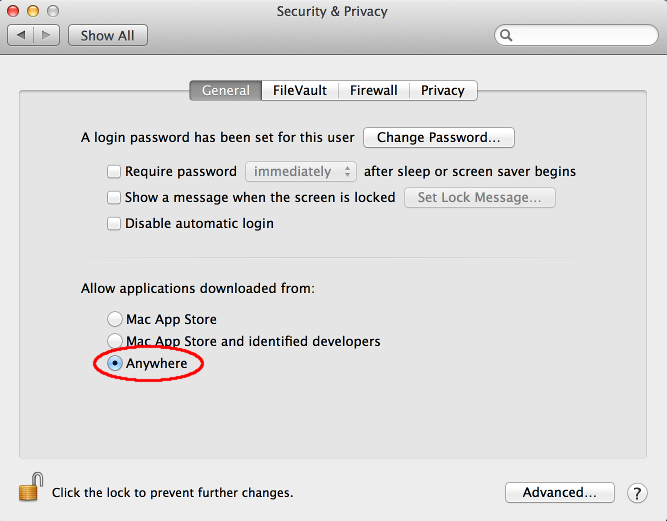
On some recent versions of Mac OS X, an error message appears ("RNAPareto is damaged and can't be opened."). This happens because Gatekeeper disallows starting unsigned apps downloaded from the internet by default. To run RNAPareto, apply the following steps:

The software is written in Java 6 and ISO C++11. To run, a Java Runtime Environment, version ≥1.6.0 is required. It is well tested with GCC 4.6. On MAC, GCC 4.2 is sufficient.
This program is free software; you can redistribute it and/or modify it under the terms of the GNU General Public License (GPL) 3.0.